The genetic evolution of European yew (Taxus baccata) during the Quaternary Ice Age
A summary of Mayol, et al. 2015. Adapting through glacial cycles: insights from a long-lived tree (Taxus baccata).
May 2015
In 2015 the New Phytologist published a paper in which 14 scientists from Spain, France, Italy, and Slovakia present the results of their investigations into the genetic evolution of European yew (Taxus baccata) during the Quaternary glacial cycles.
The study found evidence that yew colonized Europe from the East, and that it diverged into two groups (Western, Eastern) at the beginning of the Quaternary glaciations, c. 2.2 million years before present. The study also discovered a significant role of environmental adaptation during interglacials at the origin of genetic divergence between both groups.
Since this process may be common in other species, the study provides new research lines to explore the effect of Quaternary climatic factors on present-day patterns of genetic diversity.
The glaciations (‘Ice Ages’) and the interim warming periods (or interglacials) of the Quaternary Period (the current and most recent geologic period, 2.58 million years ago until present) have long been known as the major influence in shaping the geographical distribution of European species and their patterns of genetic structure. But their contribution of geographic isolation and/or environmental adaptation in creating population genetic divergence has so far been unexplored. This study used a long-lived tree as a model species to investigate the impact of Quaternary climatic changes on genetic diversity via neutral (isolation-by-distance, IBD) and selective (isolation-by-adaptation, IBA) processes. Trees being long-lived organisms are particularly well-suited as model species because…
• they are distributed over large areas with a wide spectrum of both biotic (e.g. plant community structure, browsing pressures) and abiotic (e.g. terrain, soil, exposure, precipitation) factors,
• they show local adaptation to environmental gradients in many respects of geography and locality,
• their life history traits (such as great longevity, overlapping generations, prolonged juvenile phase) buffer effects on changes in genetic structure.
Other studies (on oaks, Q. engelmannii, Q. lobata) had already shown that ‘the genetic signal of past climate can persist over extended time periods in organisms with large effective population sizes and long generation times.’
Materials and Methods
To assess the most likely time course for the appearance of environmental barriers to gene flow (and hence their role as actual contributors to ‘isolation-by-adaptation’, IBA), various palaeoclimatic databases could be used, testing each period separately. The recently developed approximate Bayesian computation (ABC) methods helped to elucidate complex demographic scenarios as well as to estimate their timescales. Finally, the available climatic information for three time periods –
• the last interglacial (LIG, c. 120,000–140,000 yrs before present, BP),
• the last glacial maximum (LGM, c. 21,000 yrs BP), and
• present conditions (PRE, c. 1950–2000) –
*BP = before present
was used to evaluate the relative importance of current and past climatic conditions on the observed patterns of genetic variation. Coupling these approaches helped to determine whether standing patterns of genetic divergence are the result of historical isolation or, alternatively, of local adaptation to ecologically differentiated areas.
Sampling, DNA extraction and nuclear microsatellite genotyping: A total of 4992 samples (n = 1–60 per locality, mean 21) were collected at 238 localities covering the entire distribution range of Taxus baccata L. Total DNA was isolated from 50 to 100mg of dry leaf material. Seven primer pairs for the amplification of nuclear microsatellites (nuSSRs) developed specifically for T. baccata were used for the genetic analysis.
Chloroplast DNA sequencing: Six chloroplast regions plus the trnS-trnQ spacer region were tested but only three of the regions were successfully amplified: rbcL, trnS-trnQ and trnLtrnF. The amplified products were screened for polymorphism, purified, and sequenced from both ends. Sequences available from previous studies were downloaded from Gen-Bank and aligned with the newly obtained sequences.
Genetic diversity: In all populations with at least eight individuals (195 locations, n = 4829), observed and expected heterozygosity* were computed, the number of private alleles** and allelic richness were calculated and linkage disequilibrium among all pairs of loci within each population was assessed.
*Most multicellular organisms have two sets of chromosomes, i.e. they are diploid. These chromosomes are referred to as homologous chromosomes. If both alleles at a locus (or gene) on the homologous chromosomes are the same, they and the organism are homozygous with respect to that gene. If the alleles are different, they and the organism are heterozygous with respect to that gene.
**An allele or allel is one of a number of alternative forms of the same gene or same genetic locus.
Demographic history: The ABC framework was applied to the nuSSRs data to infer the demographic history of Taxus baccata. A ‘clear westward cline of decreasing diversity from Iran to the Mediterranean area was … detected, which might be indicative of a colonization pattern’ from East to West. Therefore, four scenarios were run to test alternative hypotheses considering two or three genetic pools:
A) a ‘secondary contact’ later in time in which the western and the eastern gene pools mixed again (‘Admixed’),
B) a clear westward development (‘colonization’) in which Admixed separated from the western pool only,
C) a ‘secondary contact’ (like a), but with the separation of the eastern pool from an older pool in Iran,
D) (combined B and C, i.e.) three pools (Western, Eastern, Iranian), and Admixed having separated from the western pool only.
In other words: scenarios A and B tested for two genetic pools, C and D for three. While A and C tested Admixed as stemming from a secondary contact of the western and eastern pools, B and D tested Admixed as having developed from western alone.
A population was considered as admixed when the proportion of individuals assigned to the eastern or the western cluster was smaller than 70%.
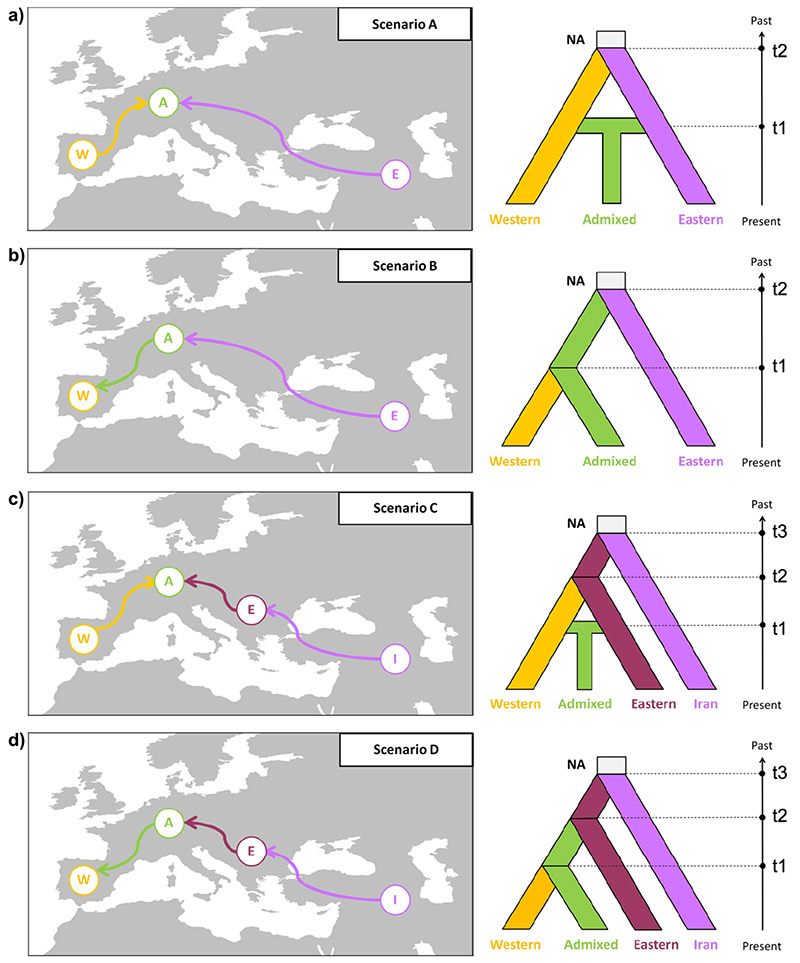
NA = ancestral effective population size; t1–t3 = divergence times for the depicted events
Since pooling data across populations can have confounding effects on the inference of demographic parameters, ten different sets of populations were constructed, each containing c. 500 individuals, representing c. 10% of the whole dataset. Each set was composed of c. 200 individuals from the Eastern pool, c. 200 individuals from the Western one and c. 100 of Admixed composition – called the ‘500-sample datasets’. For scenarios C and D, two additional datasets with four groups of populations were constructed, considering the Iran pool (Iran, Georgia) to be independent from the Eastern one. These datasets were composed by c. 200 individuals belonging to the Eastern, Western and Iran pools, respectively, and c. 100 of Admixed composition – the ‘700-sample datasets’.
One million simulations were run for each dataset.
Environmental factors: Climatic information available at the WorldClim database was applied to the locations of the same 195 yew populations to evaluate the effect of past and present climatic conditions on current genetic structure. This was performed parallel with two models: the Community Climate System Model (CCSM) and the Model for Interdisciplinary Research on Climate (MIROC).
Results
The correlation between genetic and geographic distances was highly significant, suggesting the existence of an isolation-by-distance (IBD) pattern. The genetic distances increased toward the west, indicating that eastern populations were closer to the hypothetical ‘ancestral population’.
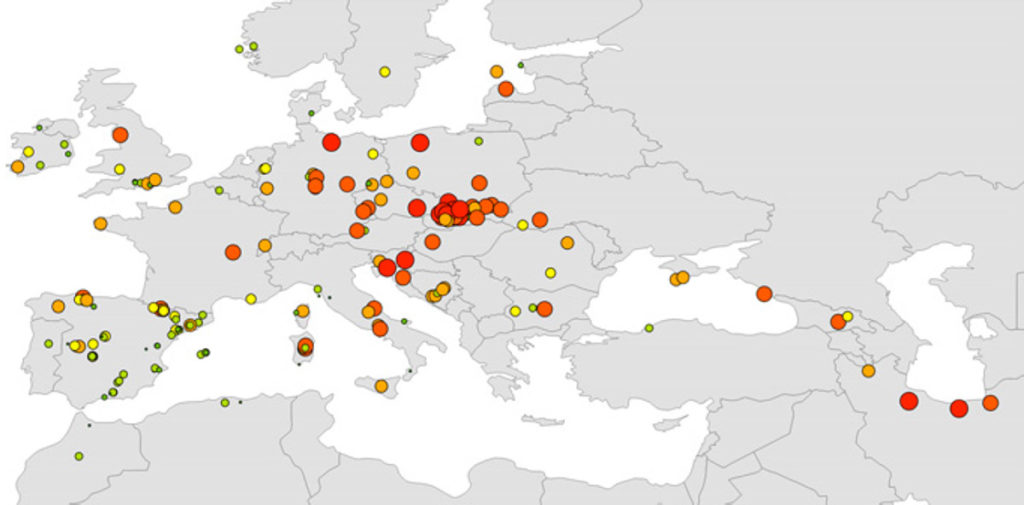
AR = allelic richness; HE = unbiased expected heterozygosity. The values for AR and HE are indicated by the circle size and the colour gradient (red > orange > yellow > green), respectively.
The most likely scenario using the ‘500-sample datasets’ was scenario A, ‘with a strong support in almost all cases’. Similarly, simulations using the ‘700-sample datasets’ unambiguously supported scenario C . Further model testing procedures endorsed the reliability of this scenario (C) as the most likely.
Under this model (scenario C), the estimated time of divergence among Iranian and European samples would have occurred about 6 million years BP* (assuming an average generation time of c. 100 years for Taxus baccata). The posterior separation of Eastern and Western clusters would have taken place about 2.2 million years BP, and the subsequent admixture event 200,000 years BP*.
*90% credible intervals: 1.35 to 14.78 million years BP
**90% credible intervals: 0.5 to 7.5 million years BP
***90% credible intervals: 50,000 to 800,000 years BP
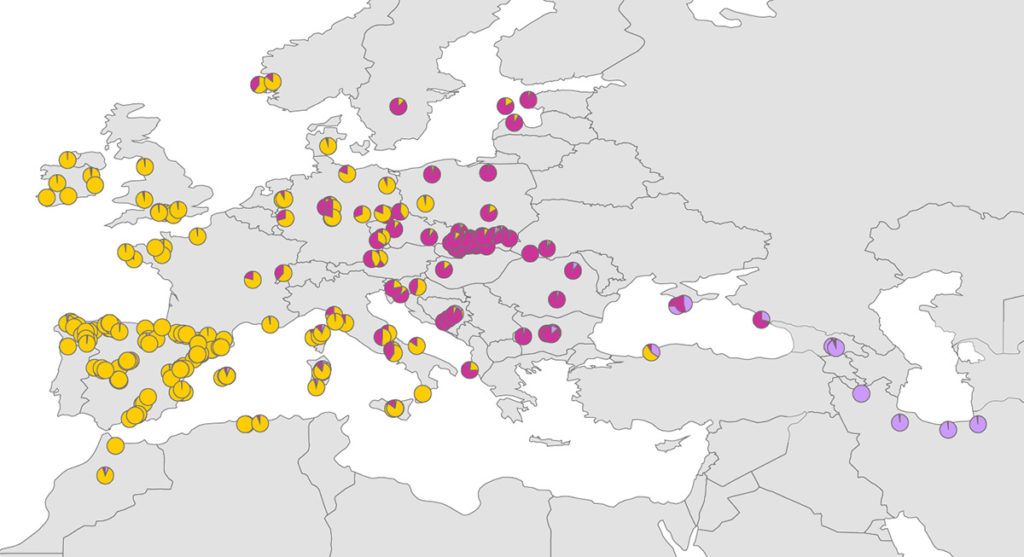
Pie charts show averaged values of the proportion of membership to each genetic cluster. Different colours indicate different genetic clusters.
The climatic investigation showed that, on average, populations within the Western cluster experienced smaller seasonal temperature fluctuations, warmer temperatures (annual mean, minimum and maximum), higher temperatures during the driest quarter, and less precipitation during the warmest quarter. There is also little overlap of their predicted distributions for all periods considered, especially during both interglacials, suggesting that Eastern and Western clusters have occupied environmentally different regions since the long past.
Discussion
The nuclear microsatellites in the genetic pool of Taxus ‘seem to retain the imprint of very ancient events’. Furthermore, ‘a near absence of variation and spatial structure for chloroplast DNA markers was observed, in accordance with the slow chloroplast nucleotide substitution rate reported in conifers’.
The ancient migration from the east is also supported by the westward decline of genetic diversity, and the westward increase in FST values (genetic distance). ‘In agreement with our results, recent studies set the origin of Taxus in North America or South West China during the late Cretaceous to middle Eocene (66.55 to 11.22 million years BP), from which it was dispersed to the current distribution areas. The genus probably reached Europe through the Irano-Turanian region, which has been postulated as a key source for the colonization of the Mediterranean region (…). This event probably occurred before the Lower Miocene, as indicated by the oldest fossil record (16–23 million years BP)’.
The ABC simulations place the separation of the Iranian and the European genetic pools at about 6 million years BP (although 90% credible intervals are wide, on average 1.35 to 14.78 Myr BP). Reduced gene flow is a reasonable assumption in Taxus, and is also suggested by high levels of pairwise genetic differentiation in this study.
‘An ancient separation is also evident from the results obtained using chloroplast DNA markers, because the only distinct haplotypes* were located at the eastern extreme of the distribution (Iran), suggesting that both groups became isolated a long time ago. This is additionally confirmed by the significantly higher number of private alleles detected at nuclear microsatellites within the Eastern pool, particularly in populations from Iran and Georgia.’
*A haplotype (short for haploid genotype) is ‘a collection of specific alleles (that is, specific DNA sequences) in a cluster of tightly-linked genes on a chromosome that are likely to be inherited together, that is, they are likely to be conserved as a sequence that survives … many generations.’ (Wikipedia)
This study supports the hypothesis that both geography and climate, particularly the interglacials, have played a significant role in shaping the genetic structure of Taxus in Europe. ‘The divergence between Western and Eastern clusters can be explained by their persistence in spatially isolated refugia during glacial periods. However, species distribution models (Fig. 4) revealed an almost nonoverlapping distribution of both groups linked to distinct climatic regimes, particularly during interglacials (…).’
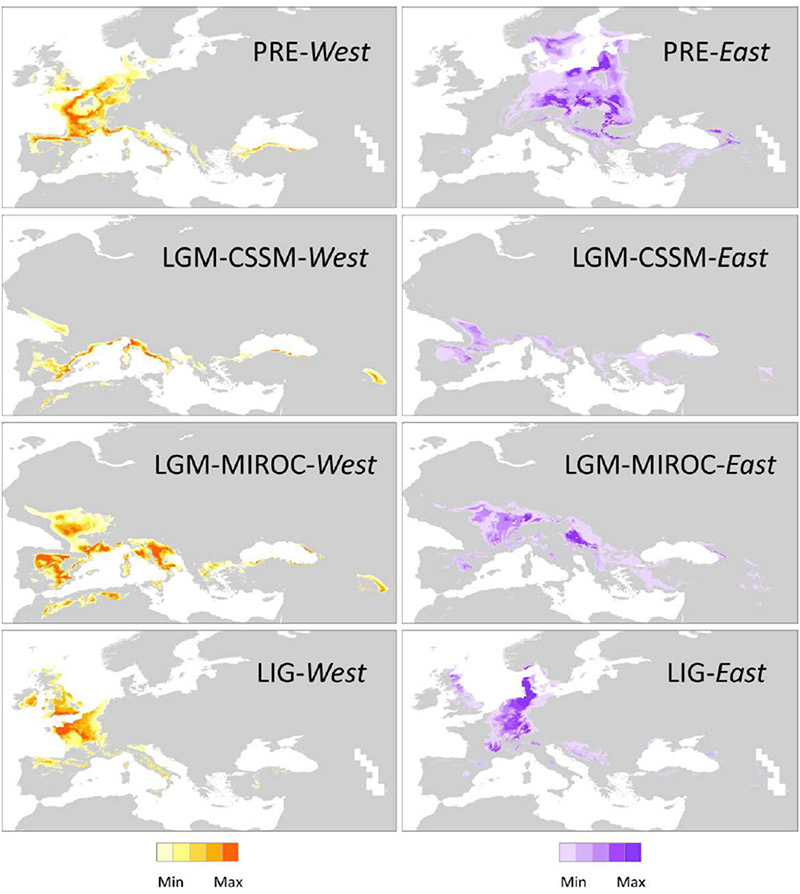
LIG, last interglacial (c. 120,000–140,000 yr before present, BP);
LGM-CCSM and LGM-MIROC, last glacial maximum (c. 21,000 yr BP) and
PRE, present conditions (c. 1950–2000).
Darker colors indicate higher probabilities of suitable climatic conditions. Areas with medium, low or no suitability are shown in grey.
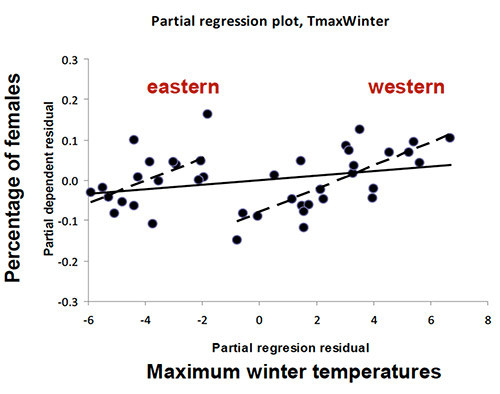
Contrary to previous ecological studies highlighting the importance of water availability on T. baccata demographic processes (such as regeneration success or population sex ratio) this study did not reveal a direct effect of rainfall on genetic divergence, but rather pointed to a major effect of temperature. There is ‘a significant association between sex-ratio and temperature, but western (Western Mediterranean and British Isles) and eastern (Central and Northern Europe) populations were clearly clustered into two distinct groups (…), suggesting the existence of two evolutionary lineages adapted to contrasted temperature ranges.’
Conclusion
‘Unravelling the exact contribution of different interglacials (Present (PRE), LIG) on environmentally driven isolation is challenging, mainly because of the absence of an extensive fossil record. Although we cannot discard temporally varying selection, there is some evidence suggesting that adaptive processes would most likely have occurred during the last interglacial.’
The results of this study provide a distinct perspective for the climatic impact of Quaternary glaciations, and how selective pressures during interglacials could have had additional impacts on genetic divergence of isolated populations. ‘This opens new lines of research to explore the effect of Quaternary climatic factors on the present-day patterns of genetic diversity in other long-lived organisms.’
Reference
Maria Mayol, Miquel Riba, Santiago C. Gonzalez-Martınez, Francesca Bagnoli, Jacques-Louis de Beaulieu, Elisa Berganzo, Concetta Burgarella, Marta Dubreuil, Diana Krajmerova, Ladislav Paule, Ivana Romsakova, Cristina Vettori, Lucie Vincenot and Giovanni G. Vendramin 2015. Adapting through glacial cycles: insights from a long-lived tree (Taxus baccata). New Phytologist (2015), doi: 10.1111/nph.13496















